Background
How do concentration index and Lorenz curves respond to different simulation scenarios?
Simulations
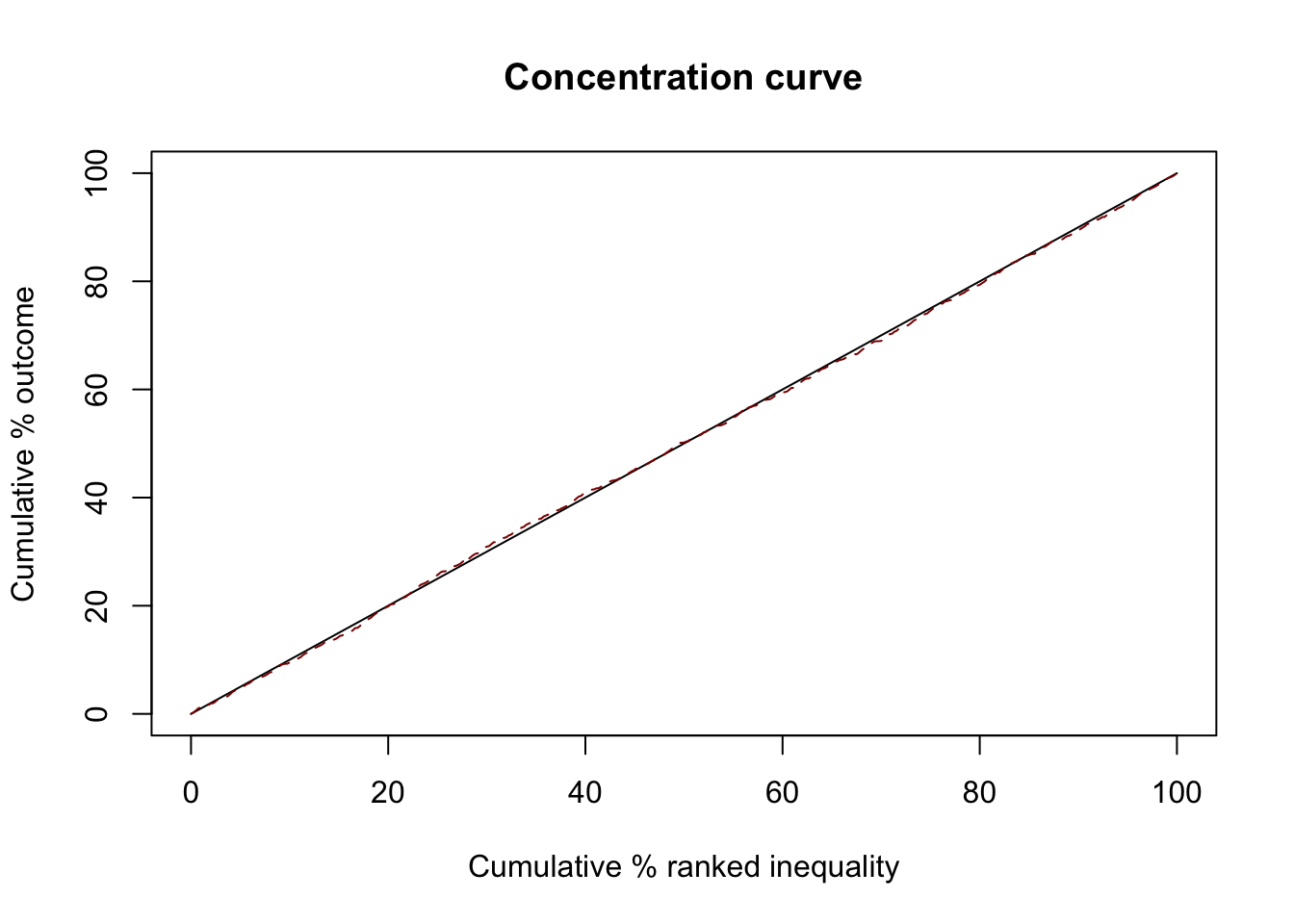
Random
GWAS attention is independent of DALY
Attaching package: 'dplyr'
The following objects are masked from 'package:stats':
filter, lag
The following objects are masked from 'package:base':
intersect, setdiff, setequal, union
library(rineq)
ndisease <- 450
diseases <- tibble(
disease = 1:ndisease,
daly = rbeta(ndisease, 1, 0.7),
gwas_attention = rbeta(ndisease, 1, 0.7))
ci1 <- ci(ineqvar=diseases$gwas_attention, outcome=diseases$daly, method="direct")
summary(ci1)
Call:
[1] "ci(ineqvar = diseases$gwas_attention, outcome = diseases$daly, "
[2] " method = \"direct\")"
Type of Concentration Index:
CI
Health Concentration Index:
0.002435111
Variance:
0.0001686933
95% Confidence Interval:
-0.02302129 0.02789151
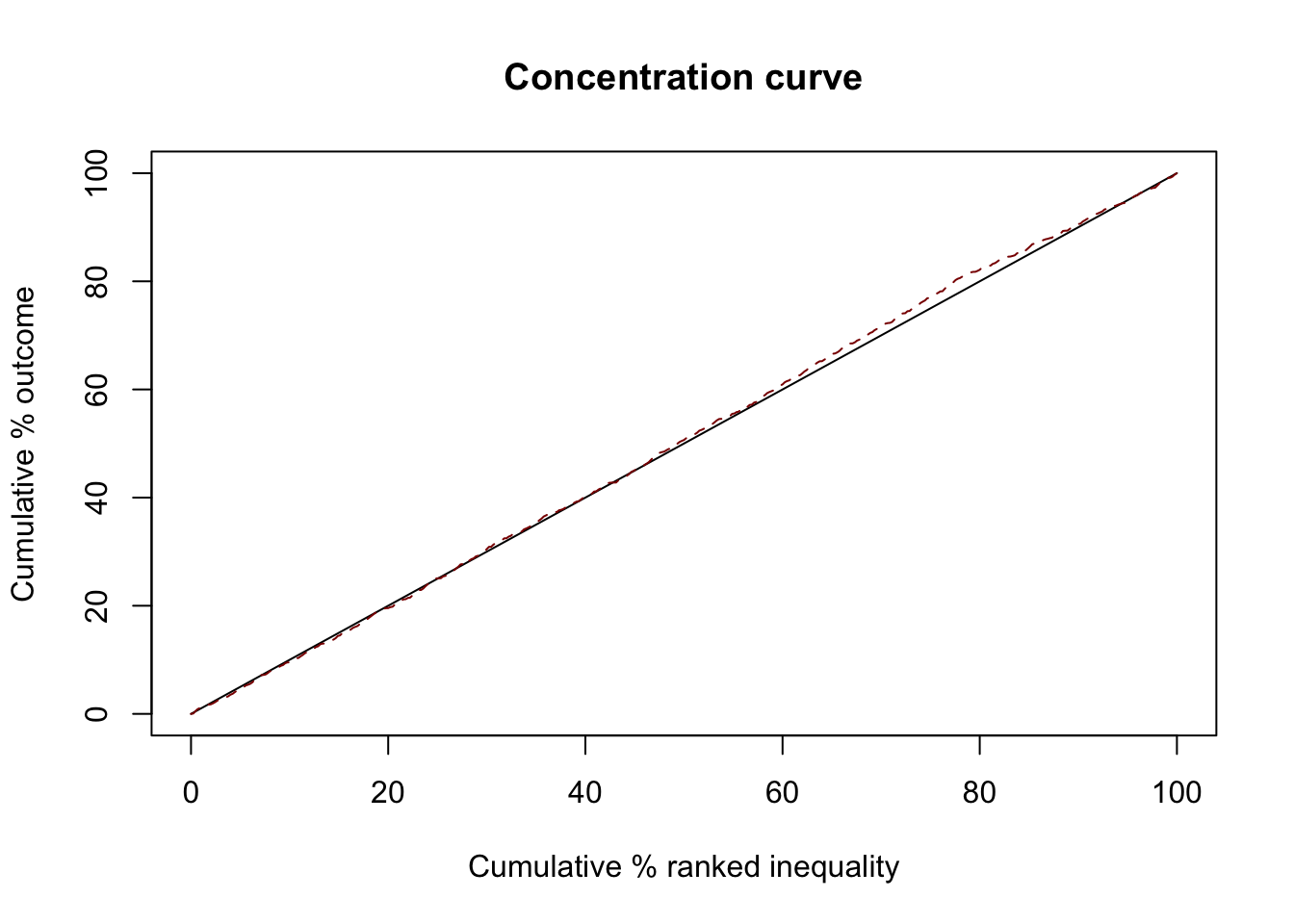
Try again with uniform distribtuion
diseases <- tibble(
disease = 1:ndisease,
daly = rbeta(ndisease, 1, 1),
gwas_attention = rbeta(ndisease, 1, 1))
ci1 <- ci(ineqvar=diseases$gwas_attention, outcome=diseases$daly, method="direct")
summary(ci1)
Call:
[1] "ci(ineqvar = diseases$gwas_attention, outcome = diseases$daly, "
[2] " method = \"direct\")"
Type of Concentration Index:
CI
Health Concentration Index:
-0.01067268
Variance:
0.0002296126
95% Confidence Interval:
-0.04037195 0.0190266
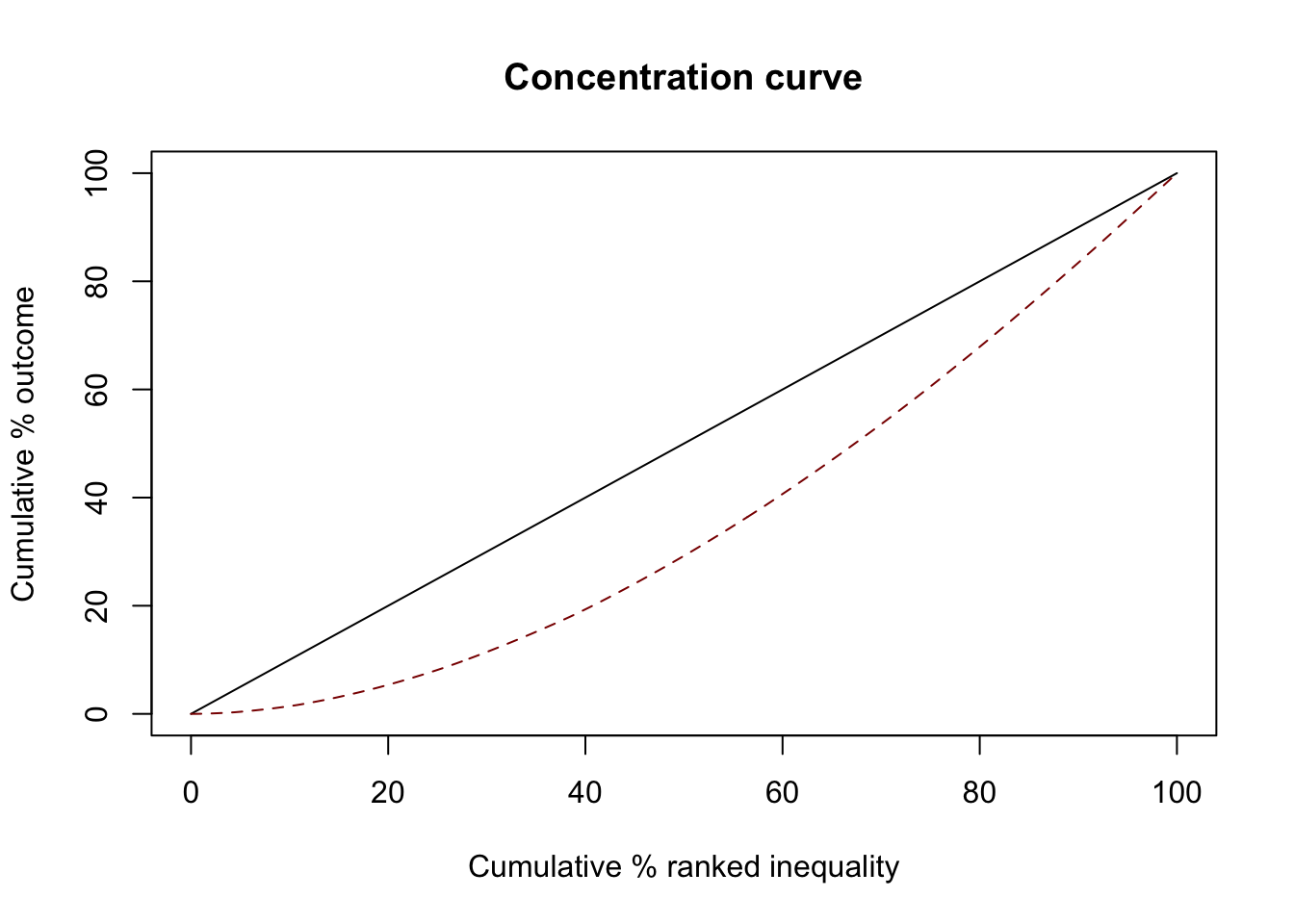
Proportional
GWAS attention is the same as DALY
ndisease <- 450
diseases <- tibble(
disease = 1:ndisease,
daly = rbeta(ndisease, 1, 0.7),
gwas_attention = daly + rnorm(ndisease, 0, sd=0.001))
ci1 <- ci(ineqvar=diseases$gwas_attention, outcome=diseases$daly, method="direct")
summary(ci1)
Call:
[1] "ci(ineqvar = diseases$gwas_attention, outcome = diseases$daly, "
[2] " method = \"direct\")"
Type of Concentration Index:
CI
Health Concentration Index:
0.277959
Variance:
0.0001193012
95% Confidence Interval:
0.2565513 0.2993668
Again with uniform distribution
diseases <- tibble(
disease = 1:ndisease,
daly = rbeta(ndisease, 1, 1),
gwas_attention = daly + rnorm(ndisease, 0, sd=0.001))
ci1 <- ci(ineqvar=diseases$gwas_attention, outcome=diseases$daly, method="direct")
summary(ci1)
Call:
[1] "ci(ineqvar = diseases$gwas_attention, outcome = diseases$daly, "
[2] " method = \"direct\")"
Type of Concentration Index:
CI
Health Concentration Index:
0.3434778
Variance:
0.0001280117
95% Confidence Interval:
0.3213023 0.3656532
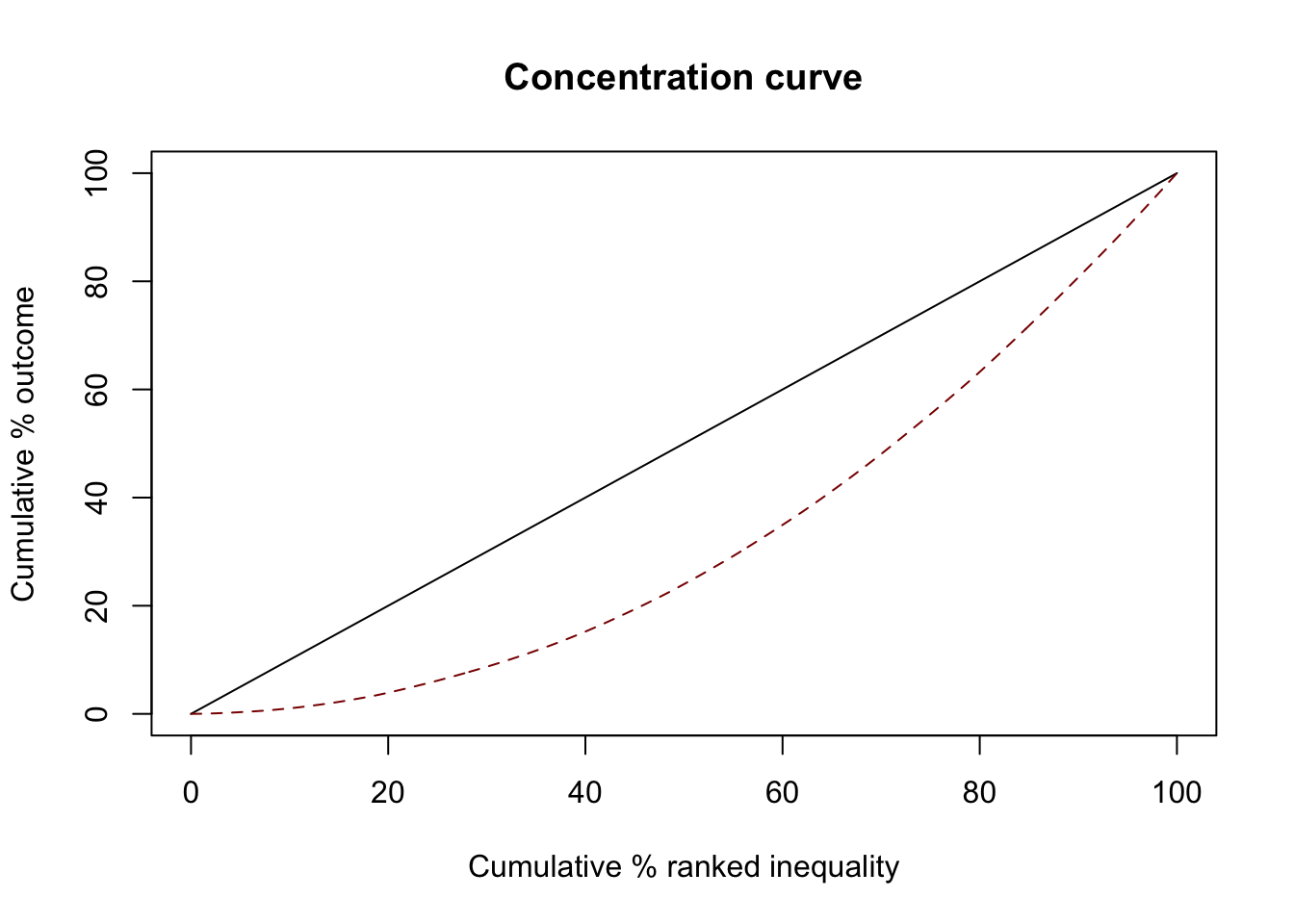
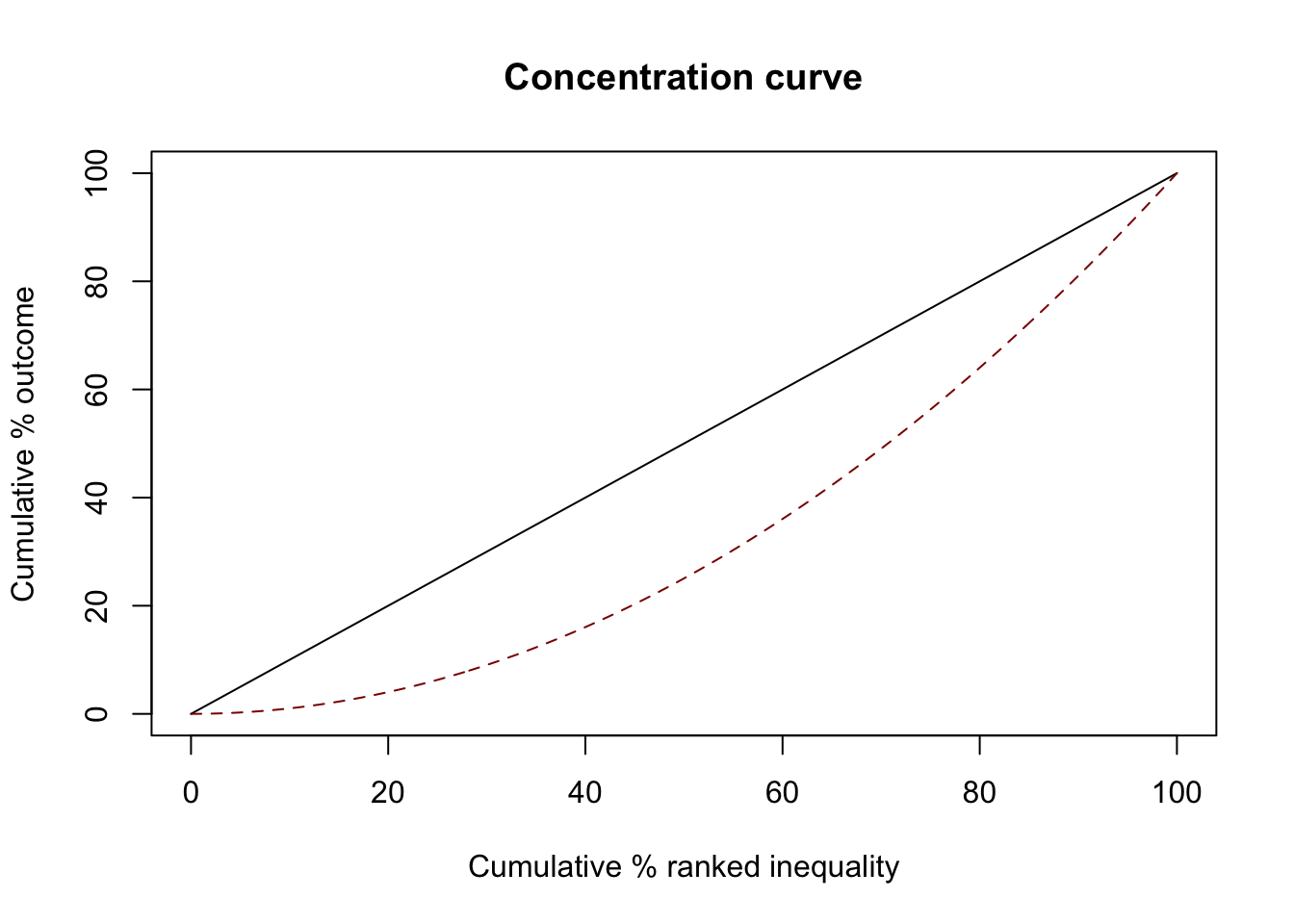
Unequal
GWAS attention grows faster than DALY
diseases <- tibble(
disease = 1:ndisease,
daly = 1:ndisease,
gwas_attention = daly^2)
ci1 <- ci(ineqvar=diseases$gwas_attention, outcome=diseases$daly, method="direct")
summary(ci1)
Call:
[1] "ci(ineqvar = diseases$gwas_attention, outcome = diseases$daly, "
[2] " method = \"direct\")"
Type of Concentration Index:
CI
Health Concentration Index:
0.3325926
Variance:
0.0001307385
95% Confidence Interval:
0.3101822 0.355003
R version 4.4.3 (2025-02-28)
Platform: aarch64-apple-darwin20
Running under: macOS Sonoma 14.6.1
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/4.4-arm64/Resources/lib/libRlapack.dylib; LAPACK version 3.12.0
locale:
[1] en_US.UTF-8/en_US.UTF-8/en_US.UTF-8/C/en_US.UTF-8/en_US.UTF-8
time zone: Europe/London
tzcode source: internal
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] rineq_0.2.3 dplyr_1.1.4
loaded via a namespace (and not attached):
[1] digest_0.6.37 utf8_1.2.4 R6_2.5.1 fastmap_1.2.0
[5] tidyselect_1.2.1 xfun_0.48 magrittr_2.0.3 glue_1.8.0
[9] tibble_3.2.1 knitr_1.48 pkgconfig_2.0.3 htmltools_0.5.8.1
[13] rmarkdown_2.27 generics_0.1.3 lifecycle_1.0.4 cli_3.6.3
[17] fansi_1.0.6 vctrs_0.6.5 compiler_4.4.3 tools_4.4.3
[21] pillar_1.9.0 evaluate_1.0.1 yaml_2.3.10 rlang_1.1.4
[25] jsonlite_1.8.9 htmlwidgets_1.6.4