# gene x drug - e.g. based on binding affinities
B <- matrix(c(
0, 1, 1,
1, 0, 0,
0, 1, 0,
0, 0, 1,
0, 0, 0,
0, 1, 1
), 6, 3)
# trait x se - matches trait names to side effect terms
D <- matrix(c(
1, 0, 0, 0, 0,
1, 0, 0, 0, 0,
0, 1, 0, 0, 0,
0, 1, 0, 0, 0,
0, 0, 1, 0, 0,
0, 0, 1, 0, 0,
0, 0, 1, 0, 0,
0, 0, 1, 0, 0,
0, 0, 0, 1, 0,
0, 0, 0, 1, 0
), 10, 5)
# True mapping of genes to side effects - we don't observe this
gse <- matrix(c(
1, 0, 0, 0, 0, 0,
1, 1, 0, 0, 0, 0,
0, 0, 1, 0, 0, 0,
0, 0, 0, 1, 1, 0,
0, 0, 0, 0, 1, 0
), 5, 6)
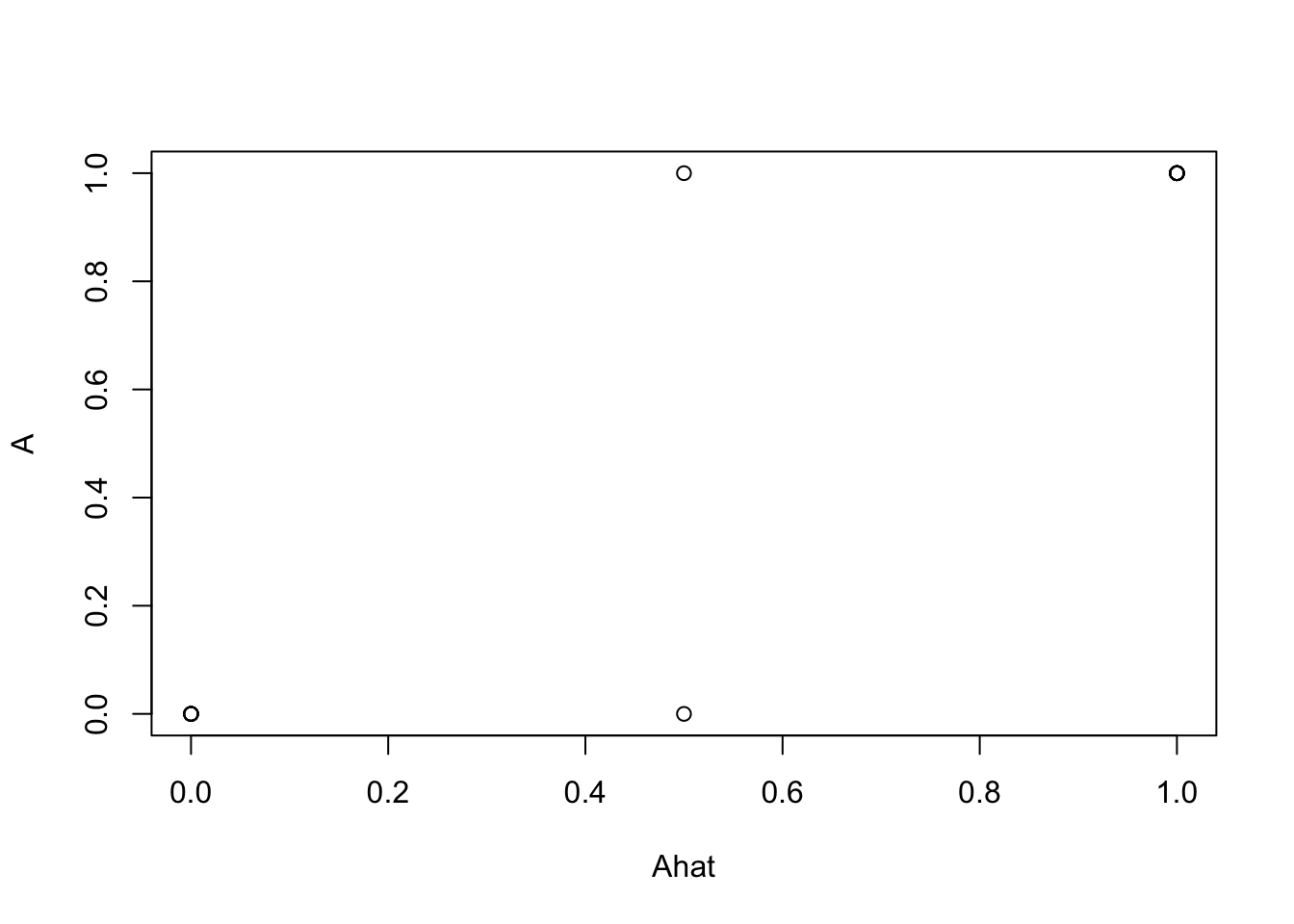
# True drug x side effect matrix is generated from gene side effects by gene drug binding
A <- gse %*% B
A