Background
Is it possible to convert BF to beta and standard error? According to Giambartolomei et al 2014 -
\[
ABF = \sqrt{1-r} \times exp(rZ^2/2)
\]
so
\[
|Z| = \sqrt{\frac{2 * log(ABF) - log(\sqrt{1-r})}{r}}
\]
here \(r = W / V\) where V is the variance of the SNP effect estimate
\[
V \approx \frac{1}{2np(1-p)}
\]
where n is sample size and p is allele frequency (assumes small amount of variance explained in trait and sd of trait is 1).
Run simulation
Use regional LD matrix to generate summary statistics with a single causal variant
Use SuSiE to perform fine mapping
Convert SuSiE Bayes Factors into Z scores, betas, standard errors
Compare converted Z, beta, se against original simulated Z, beta, SE
Simulation
Libraries
library (simulateGP)library (susieR)library (here)
here() starts at /Users/gh13047/repo/lab-book
Attaching package: 'dplyr'
The following objects are masked from 'package:stats':
filter, lag
The following objects are masked from 'package:base':
intersect, setdiff, setequal, union
Conversion function for logBF to z, beta, se
#' Convert log Bayes Factor to summary stats #' #' @param lbf p-vector of log Bayes Factors for each SNP #' @param n Overall sample size #' @param af p-vector of allele frequencies for each SNP #' @param prior_v Variance of prior distribution. SuSiE uses 50 #' #' @return tibble with lbf, af, beta, se, z <- function (lbf, n, af, prior_v= 50 )= sqrt (1 / (2 * n * af * (1 - af)))= prior_v / (prior_v + se^ 2 )= sqrt ((2 * lbf - log (sqrt (1 - r)))/ r)<- z * sereturn (tibble (lbf, af, z, beta, se))
Read in example LD matrix from simulateGP repository
<- readRDS (url ("https://github.com/explodecomputer/simulateGP/raw/master/data/ldobj_5_141345062_141478055.rds" , "rb" ))glimpse (map)
List of 3
$ ld : num [1:501, 1:501] 1 0.565 0.566 0.565 0.565 ...
..- attr(*, "dimnames")=List of 2
.. ..$ : NULL
.. ..$ : chr [1:501] "V2000" "V2001" "V2002" "V2003" ...
$ map : tibble [501 × 6] (S3: tbl_df/tbl/data.frame)
..$ chr: int [1:501] 5 5 5 5 5 5 5 5 5 5 ...
..$ snp: chr [1:501] "rs252141" "rs252140" "rs252139" "rs187544" ...
..$ pos: int [1:501] 141345062 141345192 141345218 141345361 141345678 141345805 141346830 141347360 141347465 141347931 ...
..$ alt: chr [1:501] "T" "T" "C" "G" ...
..$ ref: chr [1:501] "C" "C" "T" "T" ...
..$ af : num [1:501] 0.627 0.831 0.83 0.831 0.831 ...
..- attr(*, ".internal.selfref")=<externalptr>
$ nref: num 503
Generate summary statistics for a single causal variant and
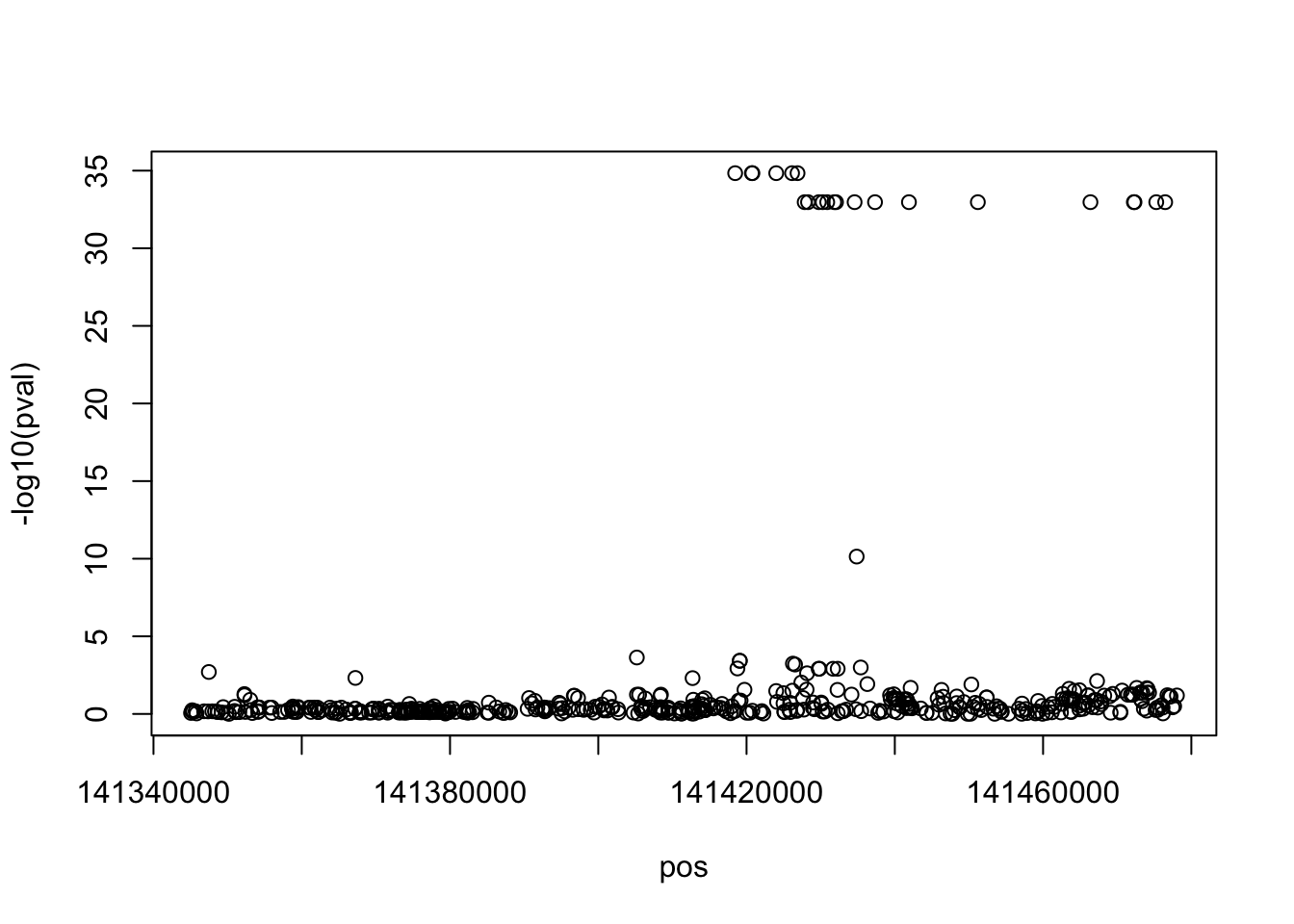
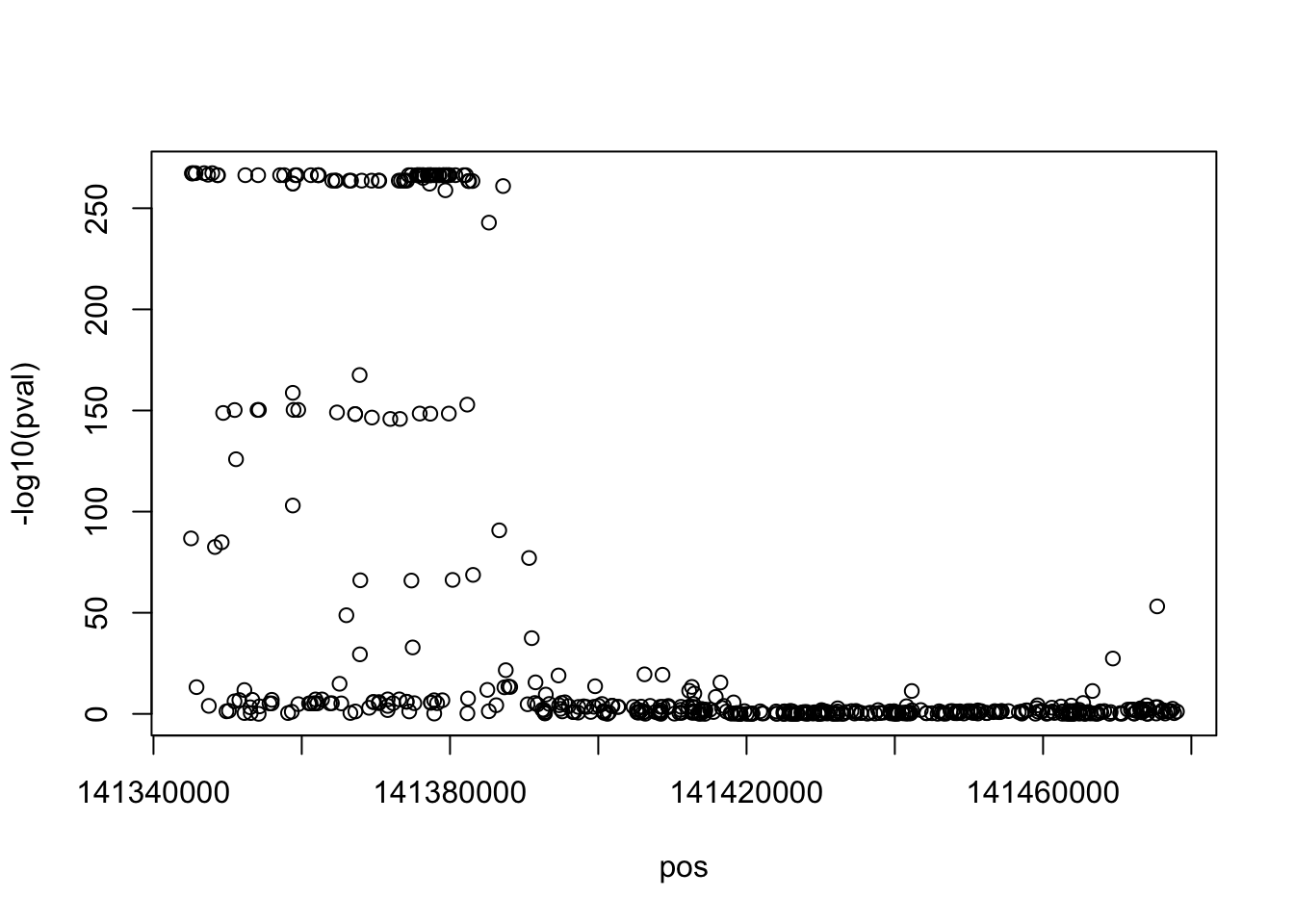
set.seed (1234 )<- map$ map %>% generate_gwas_params (h2= 0.003 , Pi= 1 / nrow (.)) %>% generate_gwas_ss (50000 , ld= map$ ld)table (ss$ beta == 0 )
plot (- log10 (pval) ~ pos, ss)
Run SuSiE
<- susie_rss (ss$ bhat / ss$ se, R = map$ ld, n = 50000 , bhat = ss$ bhat, var_y= 1 )
WARNING: XtX is not symmetric; forcing XtX to be symmetric by replacing XtX with (XtX + t(XtX))/2
Variables in credible sets:
variable variable_prob cs
286 0.1604616 1
306 0.1604616 1
291 0.1604616 1
300 0.1604616 1
274 0.1604616 1
284 0.1604616 1
Credible sets summary:
cs cs_log10bf cs_avg_r2 cs_min_r2 variable
1 30.37357 1 1 274,284,286,291,300,306
List of 18
$ alpha : num [1:10, 1:501] 6.9e-35 2.0e-03 2.0e-03 2.0e-03 2.0e-03 ...
$ mu : num [1:10, 1:501] 0.000761 0 0 0 0 ...
$ mu2 : num [1:10, 1:501] 2.04e-05 0.00 0.00 0.00 0.00 ...
$ KL : num [1:10] 6.75 -1.24e-14 -1.24e-14 -1.24e-14 -1.24e-14 ...
$ lbf : num [1:10] 6.99e+01 1.24e-14 1.24e-14 1.24e-14 1.24e-14 ...
$ lbf_variable : num [1:10, 1:501] -2.51 0 0 0 0 ...
$ sigma2 : num 1
$ V : num [1:10] 0.00307 0 0 0 0 ...
$ pi : num [1:501] 0.002 0.002 0.002 0.002 0.002 ...
$ null_index : num 0
$ XtXr : num [1:501, 1] -0.328 70.558 72.085 70.558 70.558 ...
$ converged : logi TRUE
$ elbo : num [1:2] -70876 -70876
$ niter : int 2
$ X_column_scale_factors: num [1:501] 1 1 1 1 1 1 1 1 1 1 ...
$ intercept : num NA
$ sets :List of 5
..$ cs :List of 1
.. ..$ L1: int [1:6] 274 284 286 291 300 306
..$ purity :'data.frame': 1 obs. of 3 variables:
.. ..$ min.abs.corr : num 1
.. ..$ mean.abs.corr : num 1
.. ..$ median.abs.corr: num 1
..$ cs_index : int 1
..$ coverage : num 0.963
..$ requested_coverage: num 0.95
$ pip : num [1:501] 0 0 0 0 0 0 0 0 0 0 ...
- attr(*, "class")= chr "susie"
Get Z scores from lbf
<- lbf_to_z_cont (sout$ lbf_variable[1 ,], 50000 , ss$ af, prior_v = 50 )
# A tibble: 501 × 5
lbf af z beta se
<dbl> <dbl> <dbl> <dbl> <dbl>
1 -2.51 0.373 1.41 0.00919 0.00654
2 -2.43 0.169 1.37 0.0115 0.00844
3 -2.37 0.17 1.42 0.0119 0.00842
4 -2.43 0.169 1.37 0.0115 0.00844
5 -2.43 0.169 1.37 0.0115 0.00844
6 -2.52 0.191 1.32 0.0106 0.00805
7 -2.43 0.169 1.37 0.0115 0.00844
8 -2.44 0.17 1.36 0.0115 0.00842
9 2.23 0.0139 3.17 0.0855 0.0270
10 -2.43 0.169 1.37 0.0115 0.00844
# … with 491 more rows
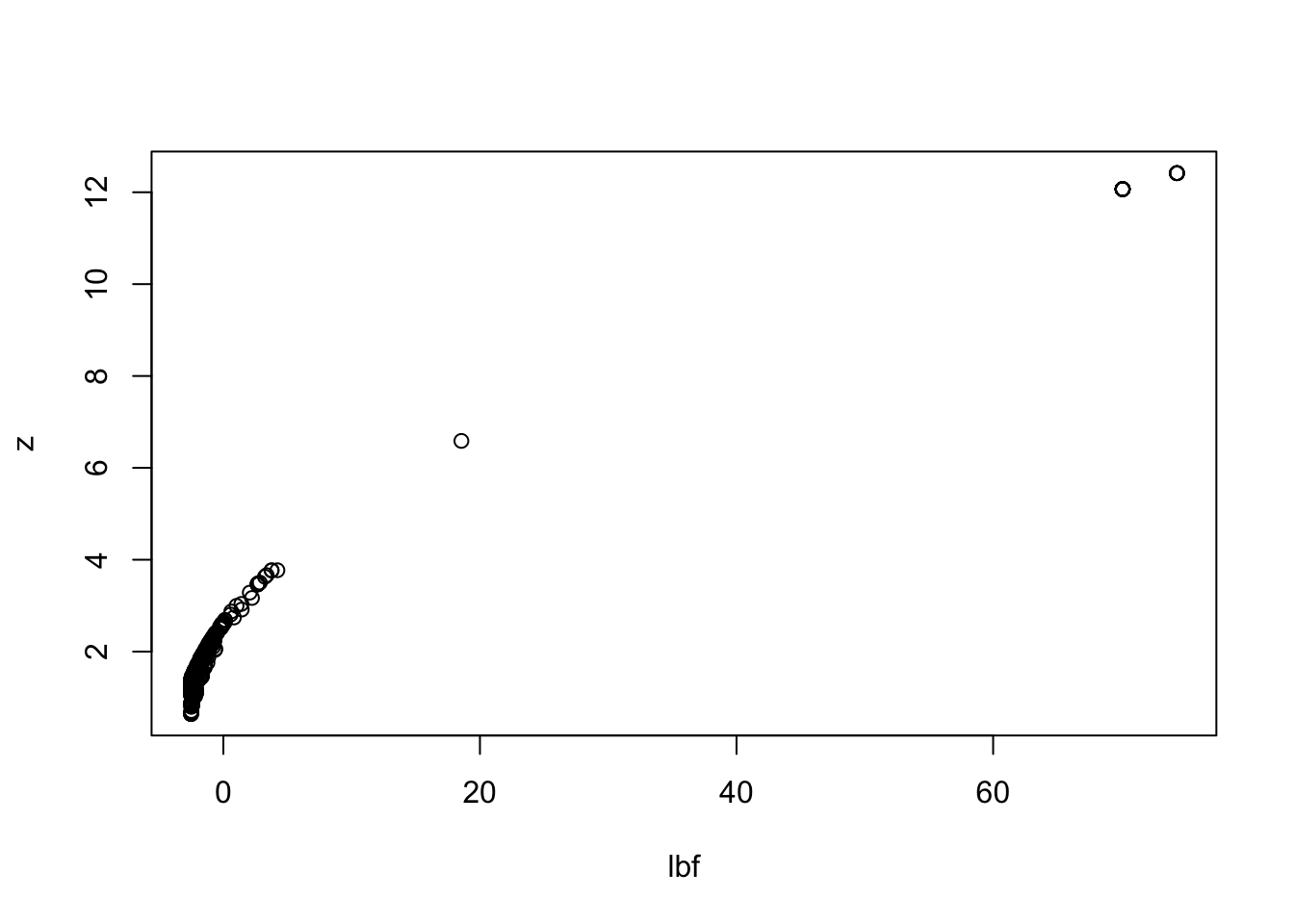
Relationship between lbf and re-estimated z
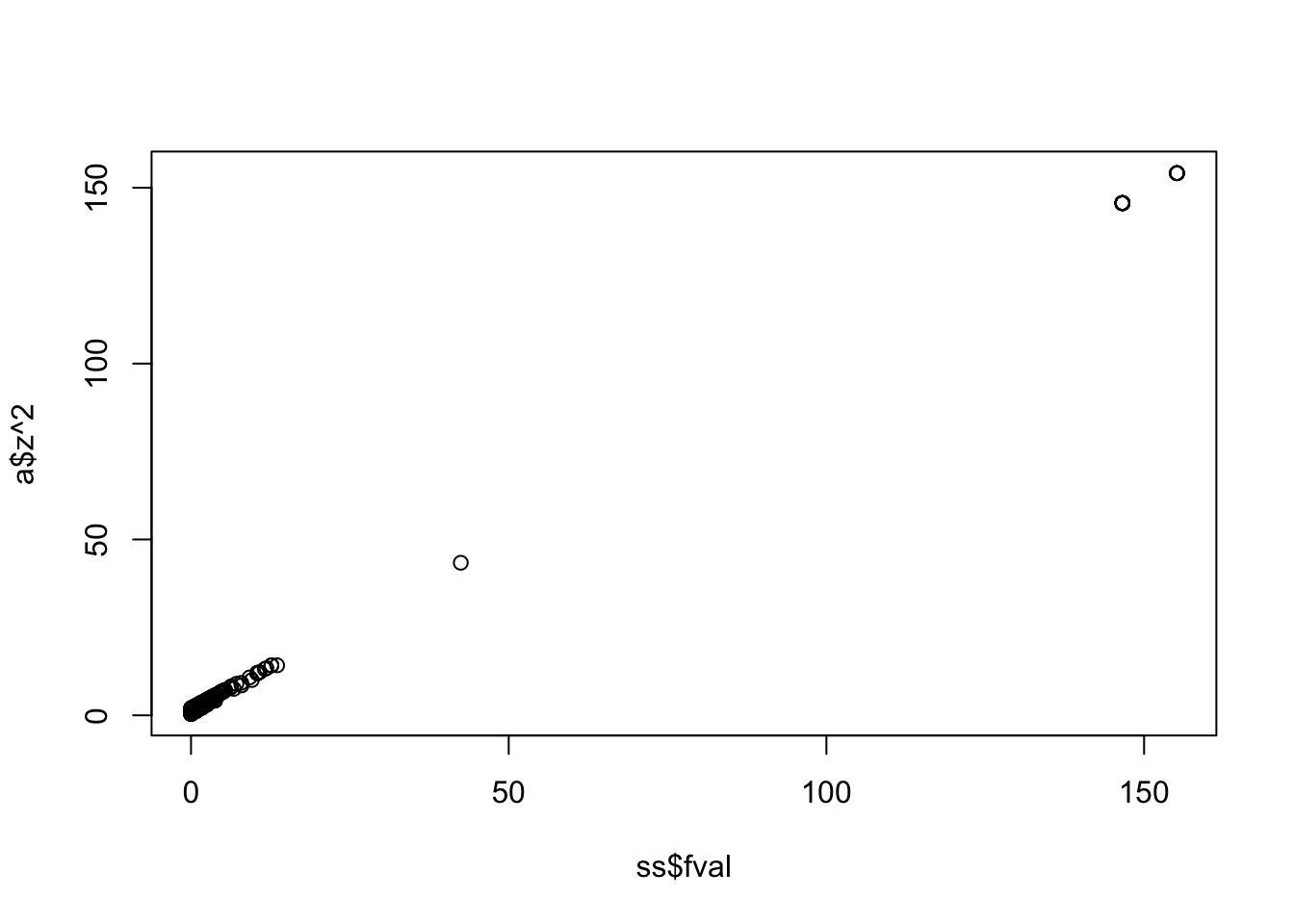
New Z vs original Z
Call:
lm(formula = a$z^2 ~ ss$fval)
Coefficients:
(Intercept) ss$fval
1.5141 0.9834
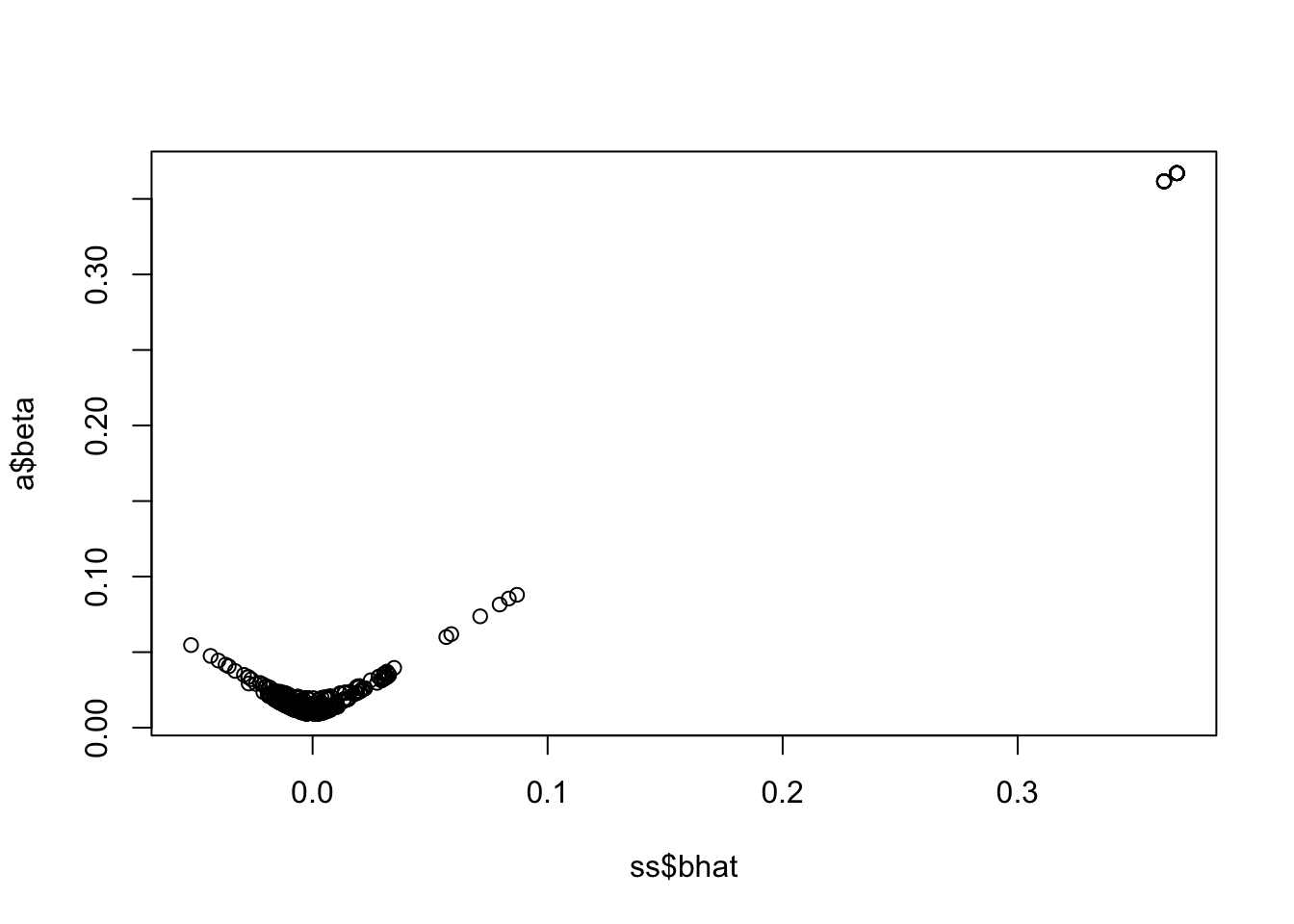
New beta vs original beta
Two causal variants
Set two causal variants at either end of the region
set.seed (12 )<- map$ map$ beta <- 0 $ beta[c (10 , 490 )] <- 0.3 <- generate_gwas_ss (param, 50000 , ld= map$ ld)plot (- log10 (pval) ~ pos, ss)
First variant
<- susie_rss (ss$ bhat / ss$ se, R = map$ ld, n = 50000 , bhat = ss$ bhat, var_y= 1 )
WARNING: XtX is not symmetric; forcing XtX to be symmetric by replacing XtX with (XtX + t(XtX))/2
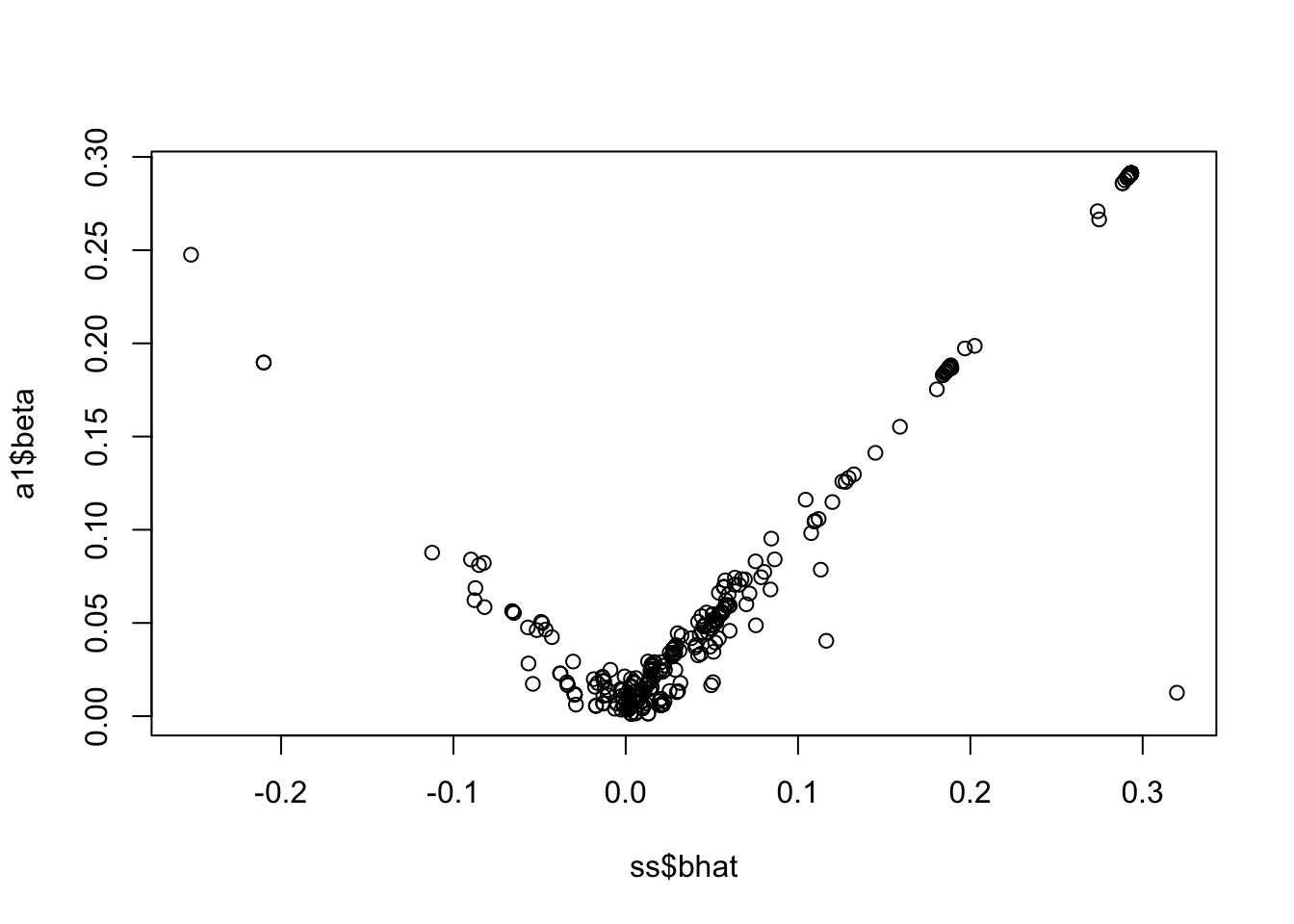
<- lbf_to_z_cont (sout$ lbf_variable[1 ,], 50000 , ss$ af, prior_v = 50 )
Warning in sqrt((2 * lbf - log(sqrt(1 - r)))/r): NaNs produced
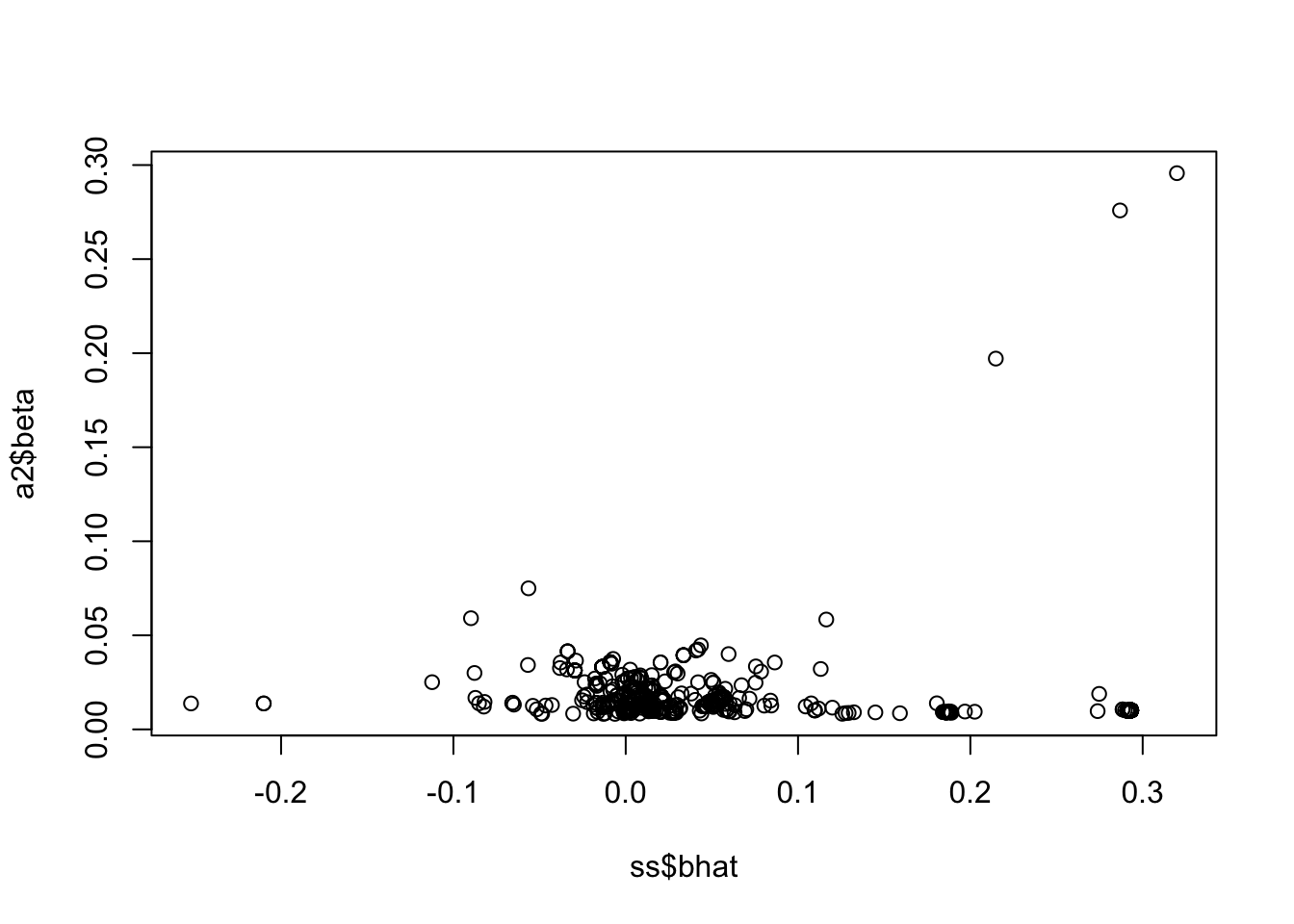
<- lbf_to_z_cont (sout$ lbf_variable[2 ,], 50000 , ss$ af, prior_v = 50 )plot (a2$ beta ~ ss$ bhat)
This looks good - it’s setting different values to 0 in the two lbf vectors that correspond to two causal variants
R version 4.2.1 Patched (2022-09-06 r82817)
Platform: aarch64-apple-darwin20 (64-bit)
Running under: macOS Monterey 12.6.2
Matrix products: default
BLAS: /Library/Frameworks/R.framework/Versions/4.2-arm64/Resources/lib/libRblas.0.dylib
LAPACK: /Library/Frameworks/R.framework/Versions/4.2-arm64/Resources/lib/libRlapack.dylib
locale:
[1] en_GB.UTF-8/en_GB.UTF-8/en_GB.UTF-8/C/en_GB.UTF-8/en_GB.UTF-8
attached base packages:
[1] stats graphics grDevices utils datasets methods base
other attached packages:
[1] dplyr_1.0.10 here_1.0.1 susieR_0.12.27 simulateGP_0.1.2
loaded via a namespace (and not attached):
[1] Rcpp_1.0.9 plyr_1.8.7 compiler_4.2.1 pillar_1.8.1
[5] tools_4.2.1 digest_0.6.31 jsonlite_1.8.4 evaluate_0.19
[9] lifecycle_1.0.3 tibble_3.1.8 gtable_0.3.1 lattice_0.20-45
[13] pkgconfig_2.0.3 rlang_1.0.6 Matrix_1.4-1 DBI_1.1.3
[17] cli_3.5.0 yaml_2.3.6 xfun_0.36 fastmap_1.1.0
[21] stringr_1.5.0 knitr_1.41 generics_0.1.3 vctrs_0.5.1
[25] htmlwidgets_1.5.4 rprojroot_2.0.3 tidyselect_1.2.0 grid_4.2.1
[29] reshape_0.8.9 glue_1.6.2 R6_2.5.1 fansi_1.0.3
[33] rmarkdown_2.16 mixsqp_0.3-48 irlba_2.3.5.1 ggplot2_3.4.0
[37] magrittr_2.0.3 MASS_7.3-58.1 matrixStats_0.63.0 scales_1.2.1
[41] htmltools_0.5.4 assertthat_0.2.1 colorspace_2.0-3 utf8_1.2.2
[45] stringi_1.7.8 munsell_0.5.0 crayon_1.5.2